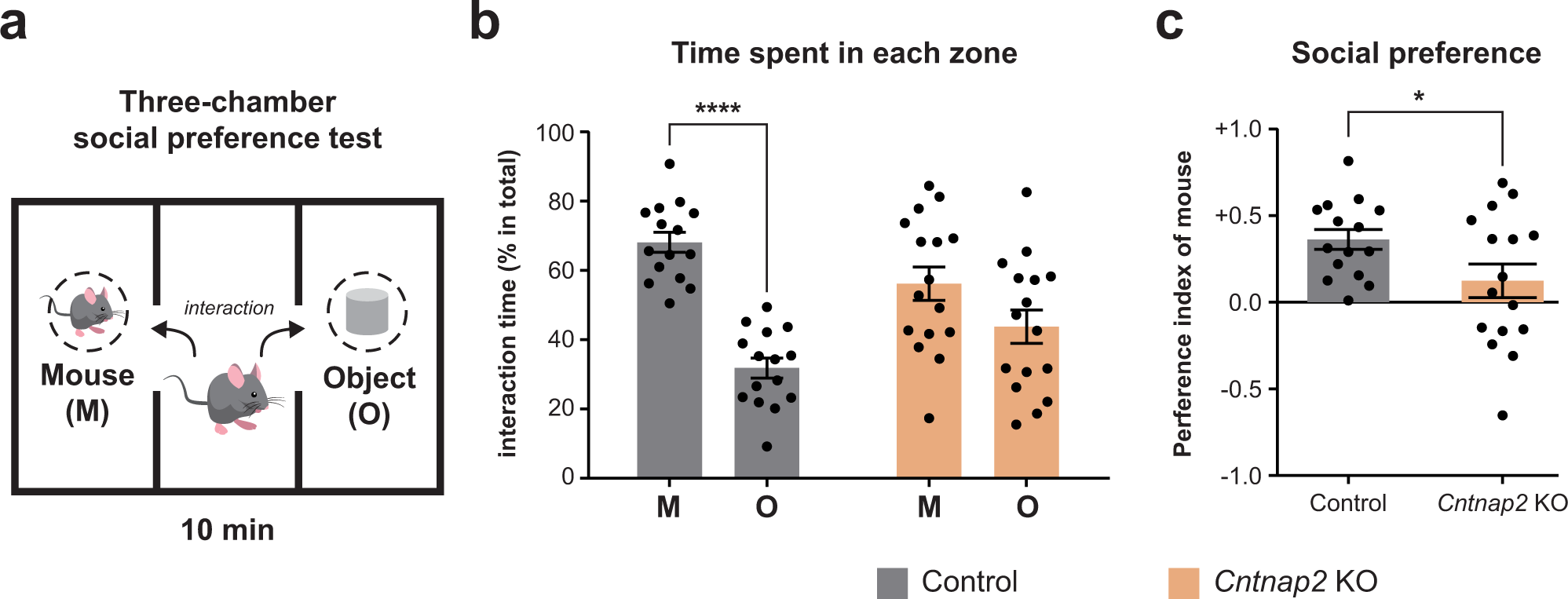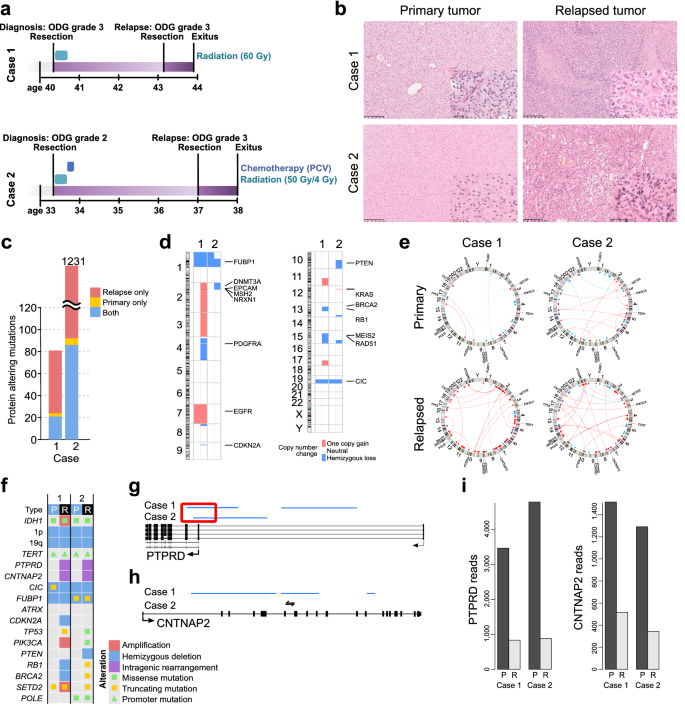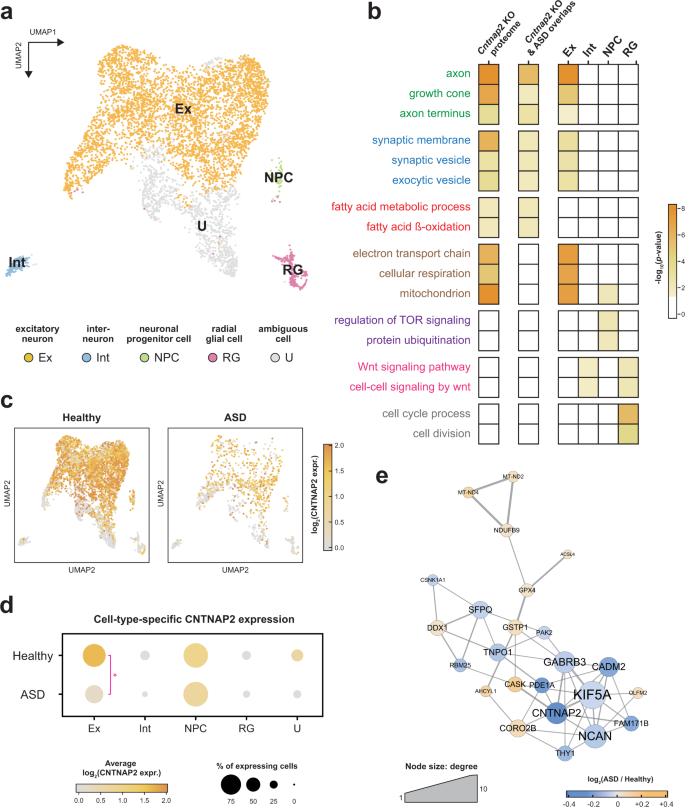
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry

Specific studies in humans and mice with ASD showing functional brain... | Download Scientific Diagram

Absence of CNTNAP2 Leads to Epilepsy, Neuronal Migration Abnormalities, and Core Autism-Related Deficits: Cell

CNTNAP2 Heterozygous Missense Variants: Risk Factors for Autism Spectrum Disorder and/or Other Pathologies? - Giorgia Canali, Laurence Goutebroze, 2018

Frontiers | Dysregulation of Parvalbumin Expression in the Cntnap2−/− Mouse Model of Autism Spectrum Disorder

Molecular Architecture of Contactin-associated Protein-like 2 (CNTNAP2) and Its Interaction with Contactin 2 (CNTN2) - ScienceDirect
No Evidence for Association of Autism with Rare Heterozygous Point Mutations in Contactin-Associated Protein-Like 2 (CNTNAP2), or in Other Contactin-Associated Proteins or Contactins | PLOS Genetics
Comprehensive cross-disorder analyses of CNTNAP2 suggest it is unlikely to be a primary risk gene for psychiatric disorders | PLOS Genetics
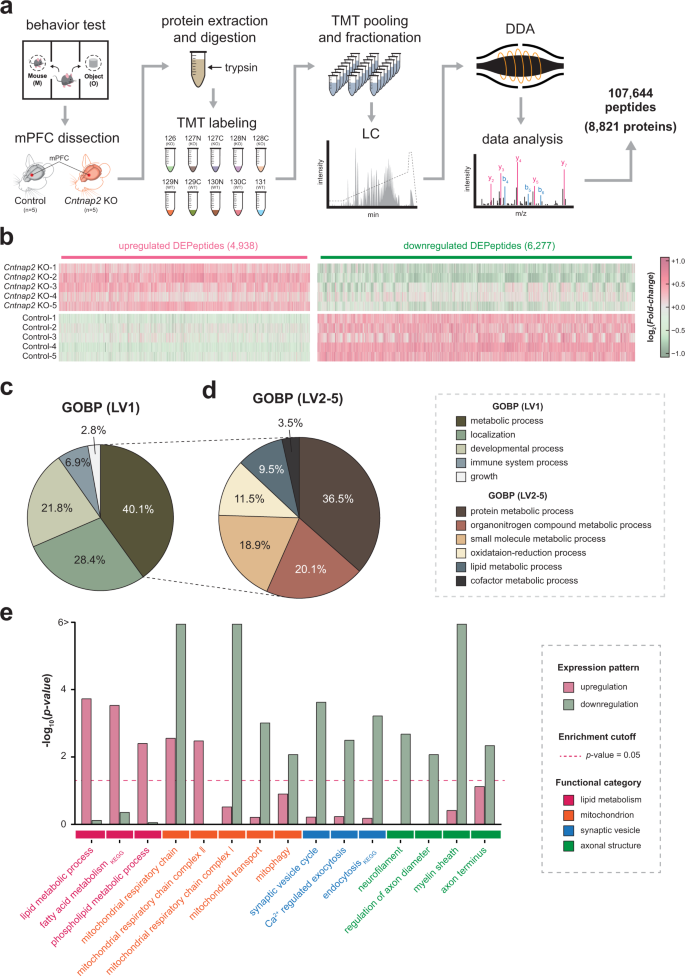
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry

PDF) Differential impacts of Cntnap2 heterozygosity and Cntnap2 null homozygosity on axon and myelinated fiber development in mouse

Exogenous and evoked oxytocin restores social behavior in the Cntnap2 mouse model of autism | Science Translational Medicine

Hippocampal gamma and sharp-wave ripple oscillations are altered in a Cntnap2 mouse model of autism spectrum disorder - ScienceDirect
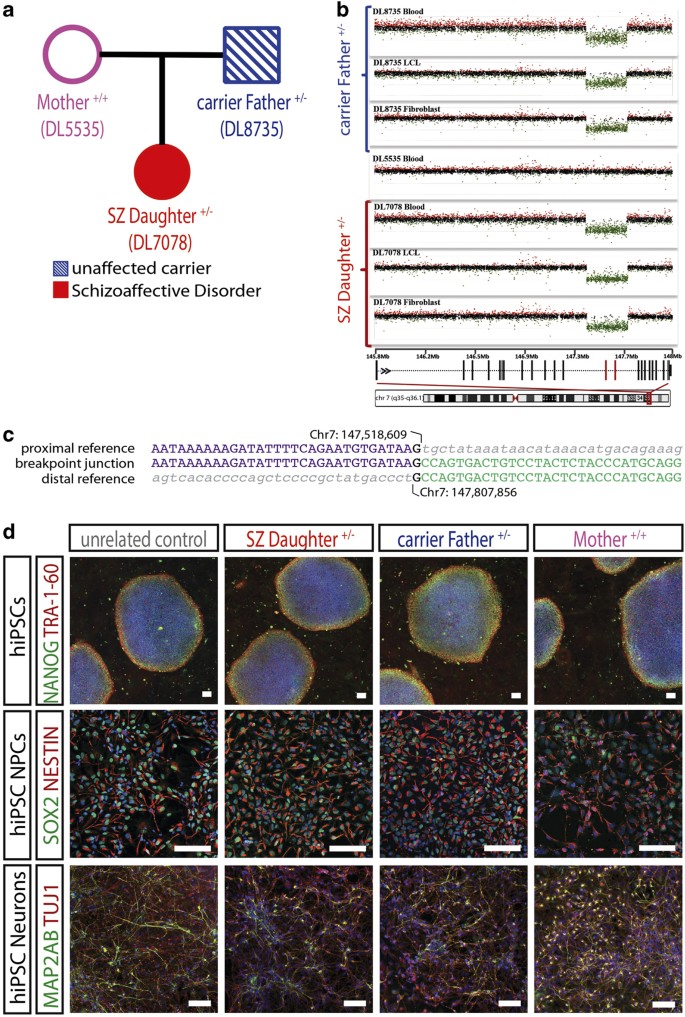
Characterization of molecular and cellular phenotypes associated with a heterozygous CNTNAP2 deletion using patient-derived hiPSC neural cells | Schizophrenia

The CNTNAP2 gene with structural rearrangements, transcription factor... | Download Scientific Diagram
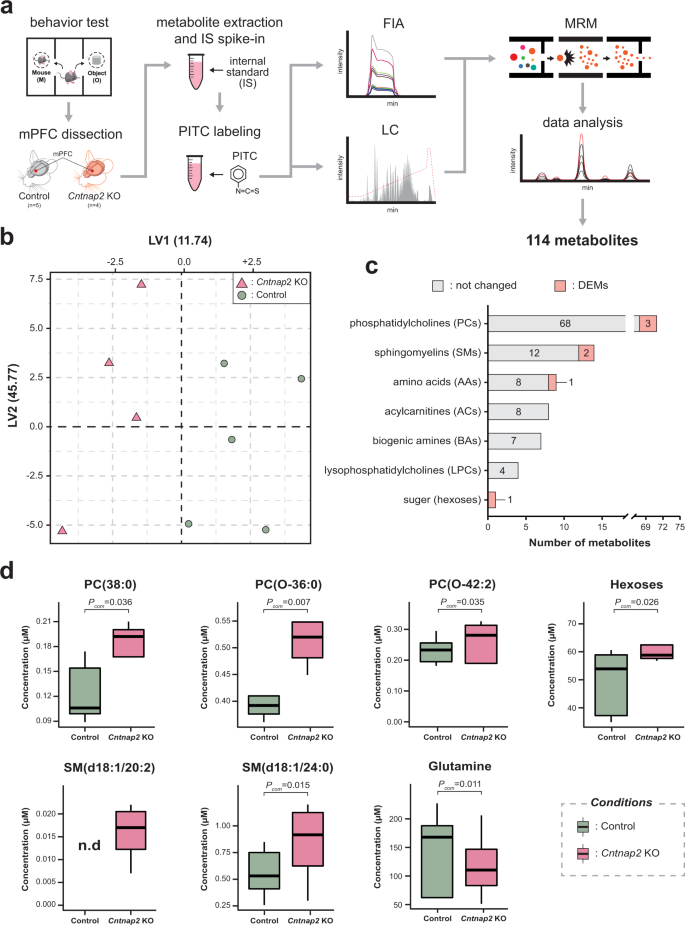
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry

JCM | Free Full-Text | Altered Blood Brain Barrier Permeability and Oxidative Stress in Cntnap2 Knockout Rat Model

CNTNAP2 Heterozygous Missense Variants: Risk Factors for Autism Spectrum Disorder and/or Other Pathologies? - Giorgia Canali, Laurence Goutebroze, 2018
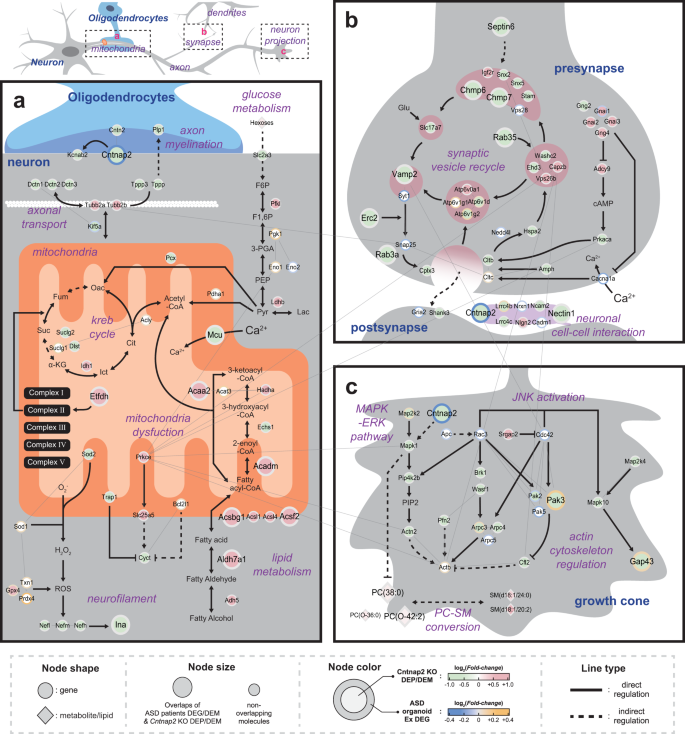
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis | Molecular Psychiatry
Comprehensive cross-disorder analyses of CNTNAP2 suggest it is unlikely to be a primary risk gene for psychiatric disorders | PLOS Genetics
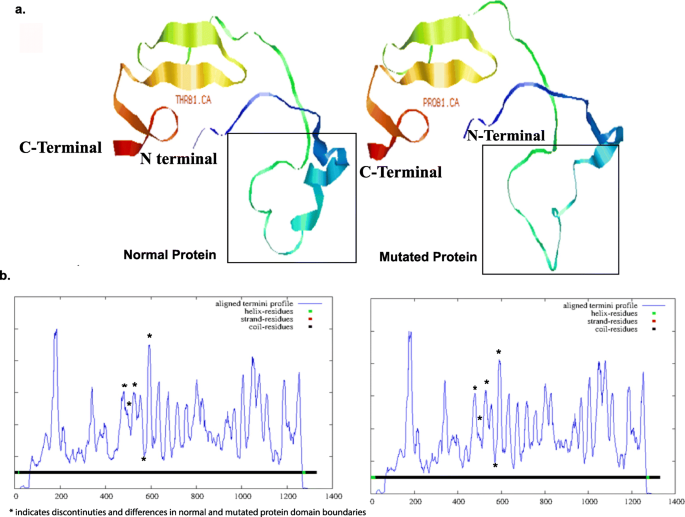
Role of CNTNAP2 in autism manifestation outlines the regulation of signaling between neurons at the synapse | Egyptian Journal of Medical Human Genetics | Full Text